Benchmarks
GRIT is benchmarked against bedtools and bedops to ensure correctness and measure performance improvements. All benchmarks verify SHA256 hash parity with bedtools output.
Summary Results
Multi-Scale Comparison (GRIT vs bedtools vs bedops)
All tools benchmarked with fair comparison settings on pre-sorted input:
| Tool | Sorted Flag | Memory Mode |
|---|---|---|
| GRIT | --assume-sorted --streaming | O(k) streaming |
| bedtools | -sorted | Streaming on sorted input |
| bedops | (requires pre-sorted) | Always streaming |
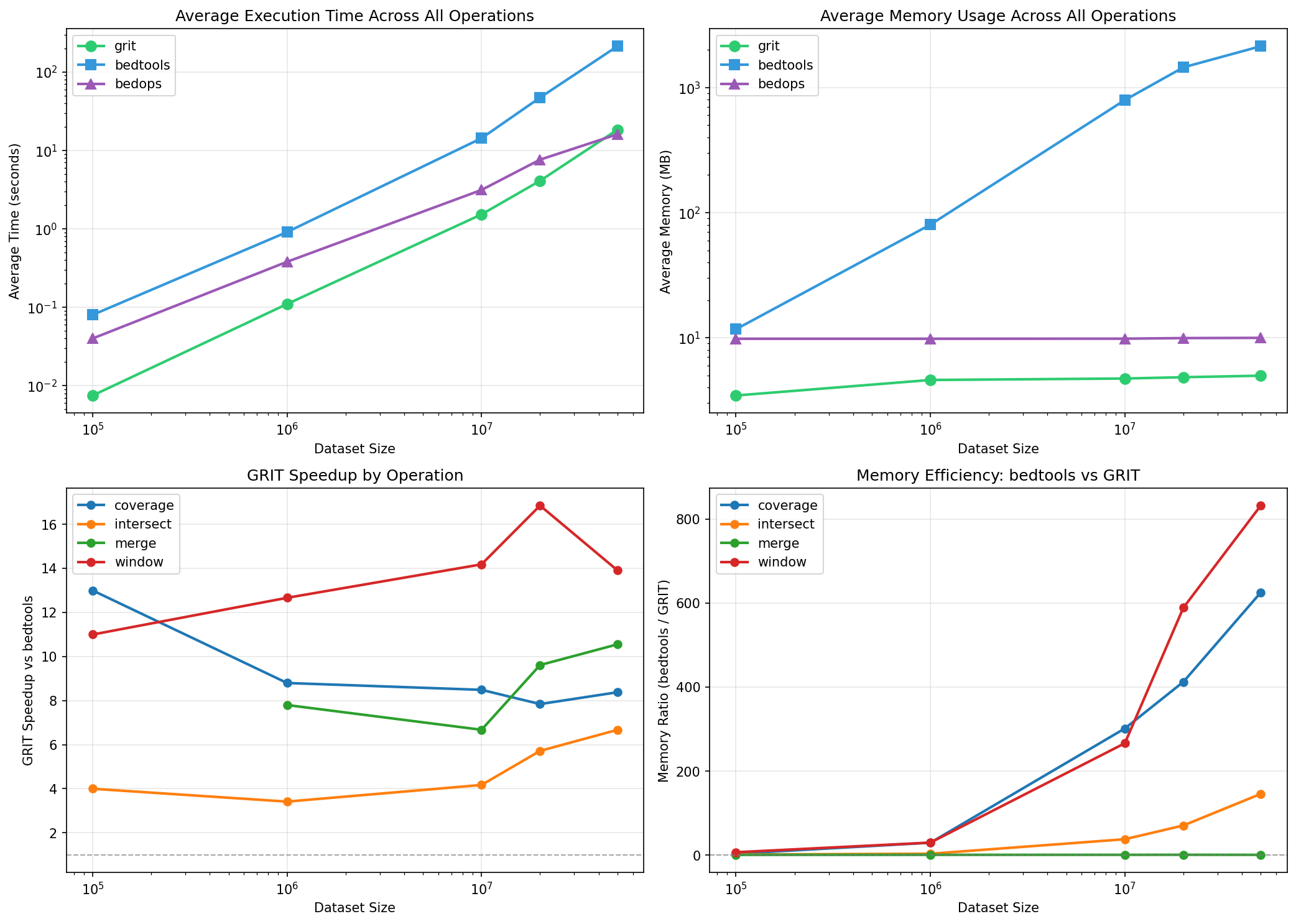
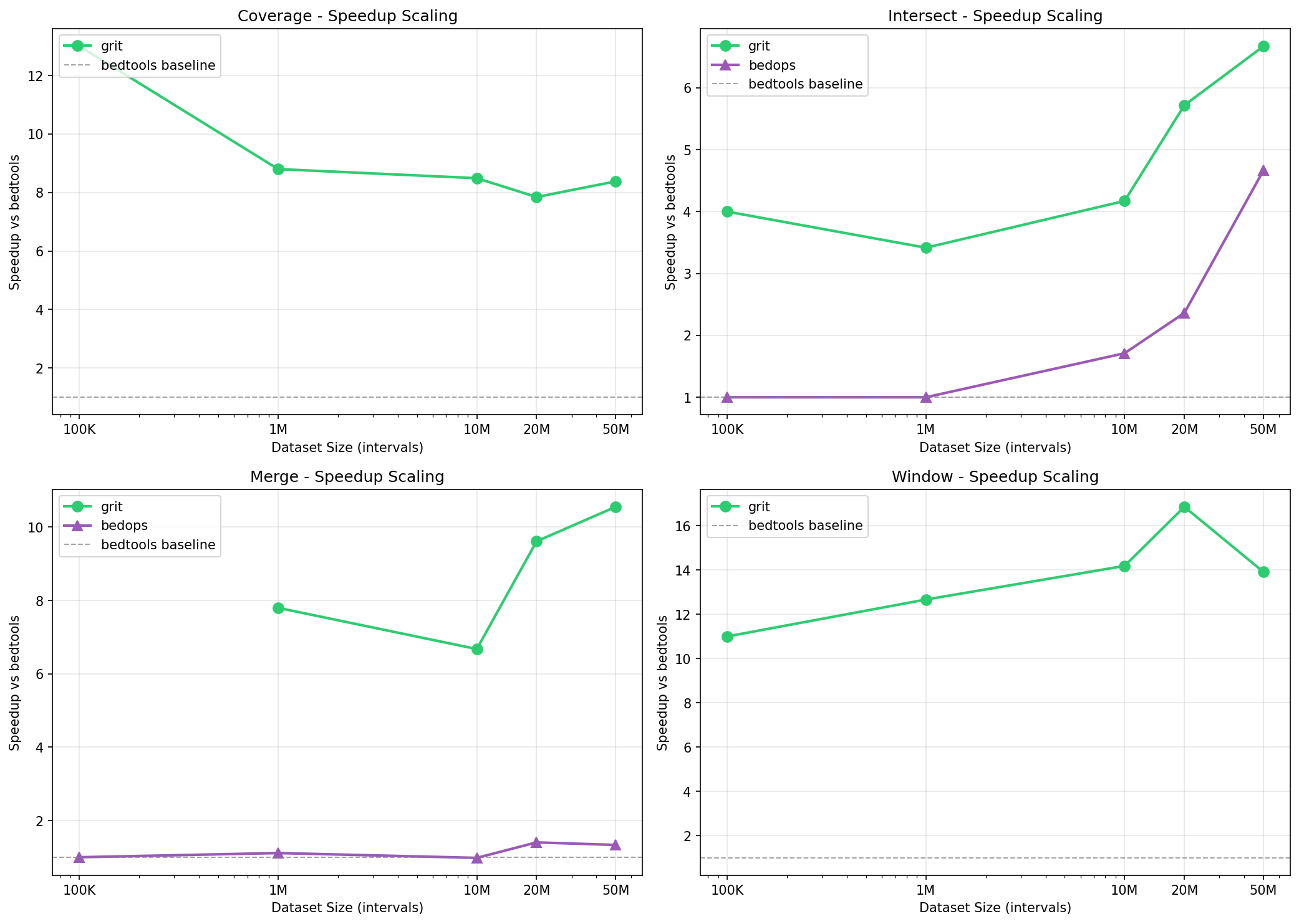
GRIT Speedup vs bedtools
| Scale | coverage | intersect | merge | window |
|---|---|---|---|---|
| 100K | 13.0x | 4.0x | - | 11.0x |
| 1M | 8.8x | 3.4x | 7.8x | 12.7x |
| 10M | 8.5x | 4.2x | 6.7x | 14.2x |
| 20M | 7.8x | 5.7x | 9.6x | 16.9x |
| 50M | 8.4x | 6.7x | 10.6x | 13.9x |
Execution Time Comparison (50M x 25M intervals)
| Command | GRIT | bedtools | bedops | GRIT Speedup |
|---|---|---|---|---|
| window | 46.5s | 10m46s | - | 13.9x |
| merge | 1.5s | 15.6s | 11.7s | 10.6x |
| coverage | 11.3s | 1m34s | - | 8.4x |
| intersect | 14.3s | 1m35s | 20.4s | 6.7x |
Memory Comparison (50M x 25M intervals)
| Command | GRIT | bedtools | bedops | Memory Savings |
|---|---|---|---|---|
| window | 5.6 MB | 4.6 GB | - | 830x less |
| coverage | 4.9 MB | 3.0 GB | - | 622x less |
| intersect | 6.0 MB | 880 MB | 10 MB | 147x less |
| merge | 3.4 MB | 3.0 MB | 10 MB | ~same |
Scaling Analysis
Scaling Charts
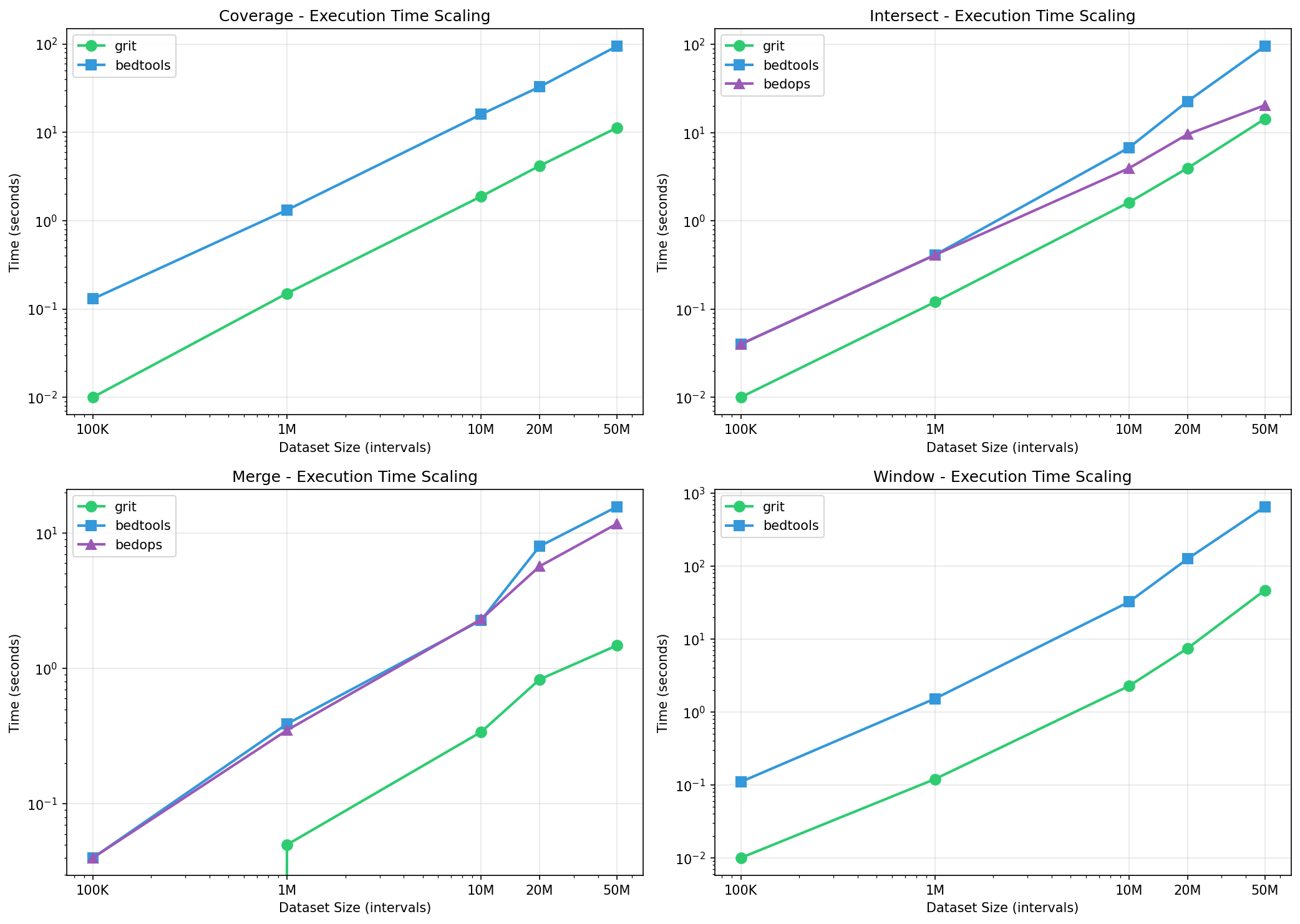
Time Scaling (Log-Log)
GRIT maintains consistent speedup advantage across all dataset sizes:
Intersect:
| Scale | GRIT | bedtools | bedops |
|---|---|---|---|
| 100K | 0.01s | 0.04s | 0.04s |
| 1M | 0.12s | 0.41s | 0.41s |
| 10M | 1.62s | 6.76s | 3.95s |
| 20M | 3.95s | 22.59s | 9.56s |
| 50M | 14.27s | 95.22s | 20.41s |
Coverage:
| Scale | GRIT | bedtools |
|---|---|---|
| 100K | 0.01s | 0.13s |
| 1M | 0.15s | 1.32s |
| 10M | 1.88s | 15.96s |
| 20M | 4.17s | 32.71s |
| 50M | 11.26s | 94.36s |
Window:
| Scale | GRIT | bedtools |
|---|---|---|
| 100K | 0.01s | 0.11s |
| 1M | 0.12s | 1.52s |
| 10M | 2.27s | 32.20s |
| 20M | 7.43s | 2m5s |
| 50M | 46.45s | 10m46s |
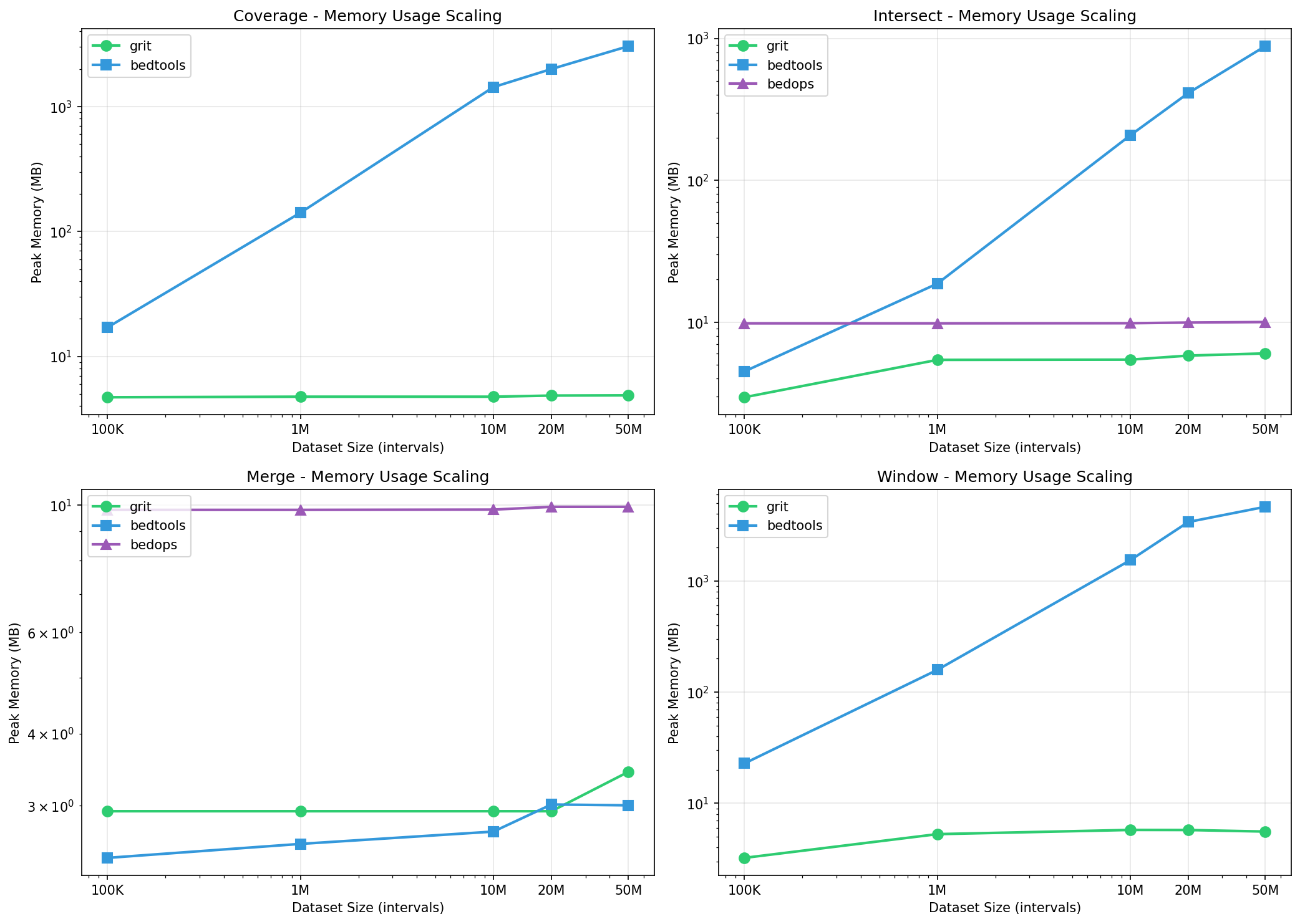
Memory Scaling
GRIT’s O(k) streaming maintains constant memory regardless of dataset size:
Coverage Memory (MB):
| Scale | GRIT | bedtools |
|---|---|---|
| 100K | 4.7 | 17 |
| 1M | 4.8 | 141 |
| 10M | 4.8 | 1,433 |
| 20M | 4.8 | 2,001 |
| 50M | 4.9 | 3,046 |
Window Memory (MB):
| Scale | GRIT | bedtools |
|---|---|---|
| 100K | 3.2 | 23 |
| 1M | 5.3 | 158 |
| 10M | 5.8 | 1,536 |
| 20M | 5.8 | 3,388 |
| 50M | 5.6 | 4,636 |
Key Observations
Memory Efficiency
GRIT’s streaming algorithms use O(k) memory where k is the maximum number of overlapping intervals at any position (typically < 100). This enables:
- Processing 50GB+ files on machines with 4GB RAM
- Constant memory regardless of file size
- No memory spikes during processing
At 50M intervals, GRIT uses 5 MB while bedtools requires 4.6 GB - an 830x reduction.
Performance vs bedops
GRIT is faster than bedops for supported operations:
- intersect: GRIT 1.4x faster than bedops at 50M scale
- merge: GRIT 7.9x faster than bedops at 50M scale
Note: bedops doesn’t support coverage or window operations.
Correctness Verification
All benchmarks verify correctness by comparing SHA256 hashes of sorted output:
PASS: bedtools hash == GRIT hash
FAIL: outputs differ (investigated and fixed)
For commands with non-deterministic output order (e.g., window), outputs are sorted before comparison.
Methodology
Test Data Generation
Synthetic BED files are generated using bedtools random:
# Generate data at various scales
./benchmarks/scale_benchmark.sh run 10M # 10M x 5M intervals
./benchmarks/scale_benchmark.sh run 50M # 50M x 25M intervals
Data characteristics:
- Genome: hg38 (24 chromosomes)
- A file interval length: 150 bp
- B file interval length: 500 bp
- B file size: 50% of A file size
- Files are sorted in genome order
Benchmark Execution
Each benchmark:
- Uses pre-sorted input files
- Runs bedtools with
-sortedflag where applicable - Runs GRIT with
--assume-sortedflag (and--streamingwhere available) - Runs bedops on pre-sorted input (no flag needed)
- Captures wall-clock time and peak RSS memory
- Verifies output correctness via line count comparison
Hardware
Benchmarks were run on:
- CPU: Apple M1 Max
- RAM: 32 GB
- Storage: NVMe SSD
- OS: macOS Darwin 24.5.0
Software Versions
- GRIT: 0.1.1
- bedtools: 2.31.1
- bedops: latest
- Rust: 1.75+
Reproducing Benchmarks
Prerequisites
# Install bedtools
brew install bedtools # macOS
# or
conda install -c bioconda bedtools # conda
# Install bedops
brew install bedops # macOS
# Build GRIT
cargo build --release
Running Benchmarks
# Quick test (100K intervals)
./benchmarks/scale_benchmark.sh quick
# Medium benchmark (100K, 1M, 10M)
./benchmarks/scale_benchmark.sh medium
# Full benchmark (100K to 100M)
./benchmarks/scale_benchmark.sh
# Specific scale
./benchmarks/scale_benchmark.sh run 50M
# Generate graphs from results
./benchmarks/scale_benchmark.sh plot
CSV Output
Benchmark results are saved as CSV for analysis:
benchmarks/results/scaling/<timestamp>/scaling_results.csv
Generating Visualizations
# Generate scaling graphs
python3 benchmarks/scripts/plot_scaling.py results.csv -o output_dir/
Real-World Datasets
GRIT can be benchmarked against real genomic data:
# List available datasets
./benchmarks/bench.sh real-list
# Download and prepare dbSNP data
./benchmarks/bench.sh real-download dbsnp --yes
./benchmarks/bench.sh real-prepare dbsnp
# Generate bedtools baseline
./benchmarks/bench.sh real-truth dbsnp all
# Run GRIT benchmark
./benchmarks/bench.sh real-run dbsnp all
Available datasets:
- dbsnp: dbSNP variant positions
- encode_peaks: ENCODE ChIP-seq peaks
- gencode: GENCODE gene annotations
- sv: Structural variant calls
Performance Tips
For maximum performance:
- Pre-sort files: Use
grit sortonce, then--assume-sorted - Use streaming mode:
--streamingfor large files - Adjust threads:
-t Nto control parallelism - Pipeline operations: Chain commands with pipes
# Optimal pipeline for large files
grit sort -i raw.bed | grit merge -i - --assume-sorted | grit intersect ...